Understanding Genetic Modification
In order to understand the process of gene manipulation, one must understand the structure of DNA and how it determines genetics.
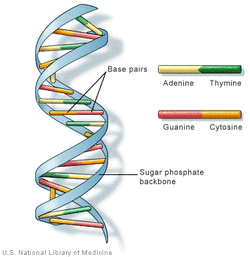
DNA stands for deoxyribonucleic acid, and is a molecule that holds the information for genetics. As seen to the right, the structure is a double helix - which is more or less like the shape of a twisted, spiral ladder - made up of two "backbone" strands of covalently bonded sugar-phosphates with nucleotide bases attached. Each nucleotide base has a specific complimentary base - adenine with thymine and guanine with cytosine - and the complimentary bases of two strands come together with strong hydrogen bonds, forming the base pairs which hold the DNA molecule together.
Genes are the traits that are determined by DNA structure. Genes can be difficult to conceptualize in the modification process because they are not items but specific traits determined by "codes" in DNA. These codes are the specific pattern of nucleotide bases in DNA. The sequence determines genetic makeup, starting with which amino acids are formed to ultimately how cells function and what traits are expressed; therefore the smallest changes to these patterns are how genes and traits are able to be modified in organisms.
In order to manipulate genes, one must know the entire genetic pattern; that is to say, the whole sequence of nucleotide base pairs must be known as well as the corresponding amino acids, so that before modifications are performed the scientist understands exactly what parts of DNA correspond to which genes. During the mechanical alteration of genes, scientists change the pattern of smaller "information packets" - segments of the genetic code that correspond to the traits they are targeting - this way the DNA will be altered in such a way that achieves the desired outcome without changing the genetic makeup of the organism completely.
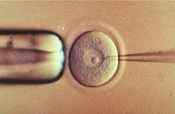
In our case study on GM cats we see this knowledge exhibited by the researchers. For this specific study, the genes of the rhesus macaque monkey were isolated and a singular gene was loaded into a lentivirus. Then the virus with the genetic information was allowed to infect an egg cell from a cat, giving the feline egg cell the new genetic information in addition to its own. This egg cell, now with augmented genetic information, was then implanted in a mother cat which would allow for the egg to grow and the genes in the initial egg to be implemented in the shaping of the new organism - thus forming the GM cat with the rhesus macaque gene (1).
DNA stands for deoxyribonucleic acid, and is a molecule that holds the information for genetics. As seen to the right, the structure is a double helix - which is more or less like the shape of a twisted, spiral ladder - made up of two "backbone" strands of covalently bonded sugar-phosphates with nucleotide bases attached. Each nucleotide base has a specific complimentary base - adenine with thymine and guanine with cytosine - and the complimentary bases of two strands come together with strong hydrogen bonds, forming the base pairs which hold the DNA molecule together.
Genes are the traits that are determined by DNA structure. Genes can be difficult to conceptualize in the modification process because they are not items but specific traits determined by "codes" in DNA. These codes are the specific pattern of nucleotide bases in DNA. The sequence determines genetic makeup, starting with which amino acids are formed to ultimately how cells function and what traits are expressed; therefore the smallest changes to these patterns are how genes and traits are able to be modified in organisms.
In order to manipulate genes, one must know the entire genetic pattern; that is to say, the whole sequence of nucleotide base pairs must be known as well as the corresponding amino acids, so that before modifications are performed the scientist understands exactly what parts of DNA correspond to which genes. During the mechanical alteration of genes, scientists change the pattern of smaller "information packets" - segments of the genetic code that correspond to the traits they are targeting - this way the DNA will be altered in such a way that achieves the desired outcome without changing the genetic makeup of the organism completely.
In our case study on GM cats we see this knowledge exhibited by the researchers. For this specific study, the genes of the rhesus macaque monkey were isolated and a singular gene was loaded into a lentivirus. Then the virus with the genetic information was allowed to infect an egg cell from a cat, giving the feline egg cell the new genetic information in addition to its own. This egg cell, now with augmented genetic information, was then implanted in a mother cat which would allow for the egg to grow and the genes in the initial egg to be implemented in the shaping of the new organism - thus forming the GM cat with the rhesus macaque gene (1).
Methods
There are several forms and techniques for modification of genetic material, depending on the goal of the change. Instances of DNA manipulation can fall under broad categories of genetic modification. Some are transgenic processes, meaning that foreign DNA has been introduced into an organism. This is opposed to cisgenic processes, where only DNA mixing that can realistically be considered to have a chance of occurring in the world is mixed, meaning mutations and breeding that seem far more likely and more natural ("Cisgenesis - definitions").
Insertion
DNA insertion is one technique for creating a change in genetic coding. The insertion relies on isolating a specific code and introducing it into an organism. This genetic modification technique can be done through a process called molecular cloning. The first step of which is isolating the genetic material for insertion ("Molecular Cloning").
The most simple isolation is a natural form of genetic isolation that has been brought into the lab for artificial use. Genes are modified through the use of already existing agents in the body. Mentioned earlier in the chapter were enzymes that could isolate DNA sequences from genetic material. These are utilized in breaking up particular sequences of DNA as part of the body's immune system, attacking foreign bodies at their core by dismantling their genetic code needed for reproduction.
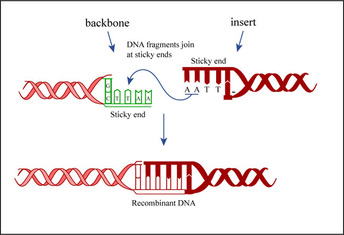
In molecular cloning, the isolated sequence, called a passenger sequence, is introduced to a sequence that codes for the reading and replication of the passenger sequence, called a vector. They are joined together by DNA ligase, which makes phosphodiester bonds between the two sequences in the ligation process. This combined sequence is recombinant DNA, rDNA. The next step in the process is to introduce the rDNA into a bacterial host, most often E. coli because of its ability to reproduce quickly. Once in the host organism, the sequence will be replicated as the organism reproduces.
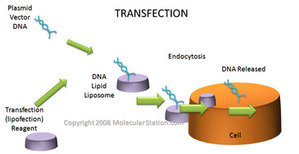
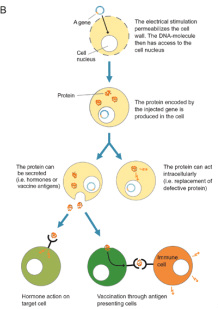
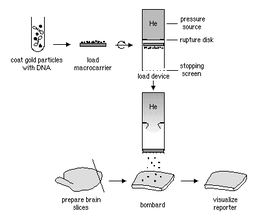
There are a variety of ways to place the rDNA into the host cell. These processes are forms of transformation. Some organisms, especially animal-celled organisms, can have genetic information inserted directly into the nucleus by microinjection, involving a needle whose end is around 1 μm in size (Evans, 2006). There are other methods as well; for instance, in the case of transfection, genetic material is planted into a virus which then implants the genetic code into organisms that it comes in contact with on a cellular level (Melcher, 2000). Transfection is what we see in the case of the GM cats. Another option is electroportation, where DNA fragments are placed in suspension with cells and a current is passed through the system. The electrical pulse disrupts the membrane of the cells and allows the DNA to enter into the cell. An additional method of introducing genetic data is biolistics. In biolistics tiny metal fragments are coated in DNA and then shot into recipient cells (Eaglesham, 2006).
Insertion
DNA insertion is one technique for creating a change in genetic coding. The insertion relies on isolating a specific code and introducing it into an organism. This genetic modification technique can be done through a process called molecular cloning. The first step of which is isolating the genetic material for insertion ("Molecular Cloning").
The most simple isolation is a natural form of genetic isolation that has been brought into the lab for artificial use. Genes are modified through the use of already existing agents in the body. Mentioned earlier in the chapter were enzymes that could isolate DNA sequences from genetic material. These are utilized in breaking up particular sequences of DNA as part of the body's immune system, attacking foreign bodies at their core by dismantling their genetic code needed for reproduction.
In molecular cloning, the isolated sequence, called a passenger sequence, is introduced to a sequence that codes for the reading and replication of the passenger sequence, called a vector. They are joined together by DNA ligase, which makes phosphodiester bonds between the two sequences in the ligation process. This combined sequence is recombinant DNA, rDNA. The next step in the process is to introduce the rDNA into a bacterial host, most often E. coli because of its ability to reproduce quickly. Once in the host organism, the sequence will be replicated as the organism reproduces.
There are a variety of ways to place the rDNA into the host cell. These processes are forms of transformation. Some organisms, especially animal-celled organisms, can have genetic information inserted directly into the nucleus by microinjection, involving a needle whose end is around 1 μm in size (Evans, 2006). There are other methods as well; for instance, in the case of transfection, genetic material is planted into a virus which then implants the genetic code into organisms that it comes in contact with on a cellular level (Melcher, 2000). Transfection is what we see in the case of the GM cats. Another option is electroportation, where DNA fragments are placed in suspension with cells and a current is passed through the system. The electrical pulse disrupts the membrane of the cells and allows the DNA to enter into the cell. An additional method of introducing genetic data is biolistics. In biolistics tiny metal fragments are coated in DNA and then shot into recipient cells (Eaglesham, 2006).
Recipient or host organisms are then selectively allowed to reproduce, which means only letting those with the rDNA reproduce through methods of selection and screening. In the selection process, factors like antibiotic sensitivity and nutrient requirements are used to disallow non-carriers from reproducing, while in screening all potential host organisms are allowed to reproduce and then are tested for the presence of the rDNA.
The DNA that is isolated, combined, and replicated can then be used. DNA copied during this process may be used for reintroduction into organisms, for nucleotide sequencing, for the manufacture of specific proteins in host organisms, or for the establishment of libraries of specific genetic code (Melcher, 2003).
As an alternative to molecular cloning, the specific genetic information needed for DNA insertion can also be isolated by artificial gene synthesis. In artificial synthesis, computers control highly controlled fixations of nucleotides to each other in order. This system is still viable to have errors and cannot be used to create chains of nucleotides that are longer than a couple hundred bases (Holmberg).
Gene Targeting
In the process of gene targeting, the goal is to remove or modify a genetic pattern in organisms. In order to do this, a "targeting construct" is used, which is a plasmid that has two strands of DNA similar to that of the targeted organism's DNA. The targeting construct is then inserted into embryonic cells of the organism. These embryonic cells are then introduced into healthy samples of the subject organism and allowed to mate. The offspring of the organism with embryonic injections will have the modified genetic code ("Transgenic mouse facility", 2011).
The DNA that is isolated, combined, and replicated can then be used. DNA copied during this process may be used for reintroduction into organisms, for nucleotide sequencing, for the manufacture of specific proteins in host organisms, or for the establishment of libraries of specific genetic code (Melcher, 2003).
As an alternative to molecular cloning, the specific genetic information needed for DNA insertion can also be isolated by artificial gene synthesis. In artificial synthesis, computers control highly controlled fixations of nucleotides to each other in order. This system is still viable to have errors and cannot be used to create chains of nucleotides that are longer than a couple hundred bases (Holmberg).
Gene Targeting
In the process of gene targeting, the goal is to remove or modify a genetic pattern in organisms. In order to do this, a "targeting construct" is used, which is a plasmid that has two strands of DNA similar to that of the targeted organism's DNA. The targeting construct is then inserted into embryonic cells of the organism. These embryonic cells are then introduced into healthy samples of the subject organism and allowed to mate. The offspring of the organism with embryonic injections will have the modified genetic code ("Transgenic mouse facility", 2011).
Gene Removal
Aside from insertion of DNA, another form of genetic modification is gene removal. The enzymes that naturally serve to cut into genetic codes and sever the bonds between nucleic acids are called nucleases. Most nucleases are of the restriction variety and are specific to coded sequences, meaning that they only "cut" through bonds at specific locations along DNA. Those that cut DNA across both strands at once find use in cutting out parts of DNA code from a sequence and only for removal. The enzymes used in DNA insertion are a specific kind of nuclease which cut unevenly, leaving "sticky" ends where, because of the base-pairing nature of nucleotides, it is highly favored that another strand of DNA will bond (Peters). There are also an array of artificial engineered nucleases like homing and zinc finger nucleases.
Aside from insertion of DNA, another form of genetic modification is gene removal. The enzymes that naturally serve to cut into genetic codes and sever the bonds between nucleic acids are called nucleases. Most nucleases are of the restriction variety and are specific to coded sequences, meaning that they only "cut" through bonds at specific locations along DNA. Those that cut DNA across both strands at once find use in cutting out parts of DNA code from a sequence and only for removal. The enzymes used in DNA insertion are a specific kind of nuclease which cut unevenly, leaving "sticky" ends where, because of the base-pairing nature of nucleotides, it is highly favored that another strand of DNA will bond (Peters). There are also an array of artificial engineered nucleases like homing and zinc finger nucleases.
References
1.) Coghlan, Andy. (11 September 2011). Glowing Transgenic Cats Could Boost AIDS Research.
NewScientist. Retrieved from:
http://www.newscientist.com/article/dn20896-glowing-transgenic-cats-could-boost-aids-research.html
NewScientist. Retrieved from:
http://www.newscientist.com/article/dn20896-glowing-transgenic-cats-could-boost-aids-research.html